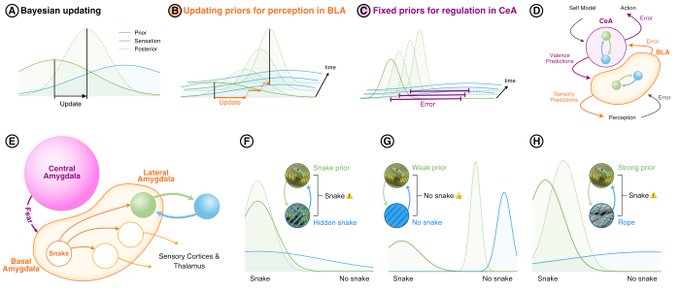
Yes! In the fMRI neurofeedback study I did in Zürich, we were able to show that people can voluntarily up- or down-regulate their amygdala. Our special twist was that we used emotional faces as feedback instead of barcharts to display the amygdala activation intensity.
In this study we asked our participants to make fearful faces less fearful or neutral faces more happy, which we presented to them while lying in our MRI scanner. We used the intensity of their amygdala activation to scale the intensity of the emotions displayed. This feasibility study worked in our healthy study population. Apurva Watve, the first author of this study, is now testing this method in people with major depression to see if it can improve their symptoms. Emotional faces could be more intuitive and naturalistic than abstract charts. We are social beings, after all.
Facing emotions: real-time fMRI-based neurofeedback using dynamic emotional faces to modulate amygdala activity
https://www.frontiersin.org/journals/neuroscience/articles/10.3389/fnins.2023.1286665/full