Hi, I'm Max Noichl! I'm a doctoral researcher working at the borders of philosophy, computational social science and digital humanities – all about doing philosophy with computers!
At the University of Bamberg, I'm working in the
SColA-project
together with Prof. Johannes Marx on a computational model for collective agency, that works under particularily adversarial condictions.
And at the University of Vienna, together with Prof. Tarja Knuuttila and Dr. Andrea Loettgers, I'm working in the
Possible Life Project on ways to computationally trace the diffusion of modeling practices through different disciplines.
When I'm not working, I play with digital art, make food, or do a little climbing. Below you can find some of my side-projects. Most are somewhat data-sciency, some also contain philosophy.
A map of presentations at EPSA23. –
Read more...
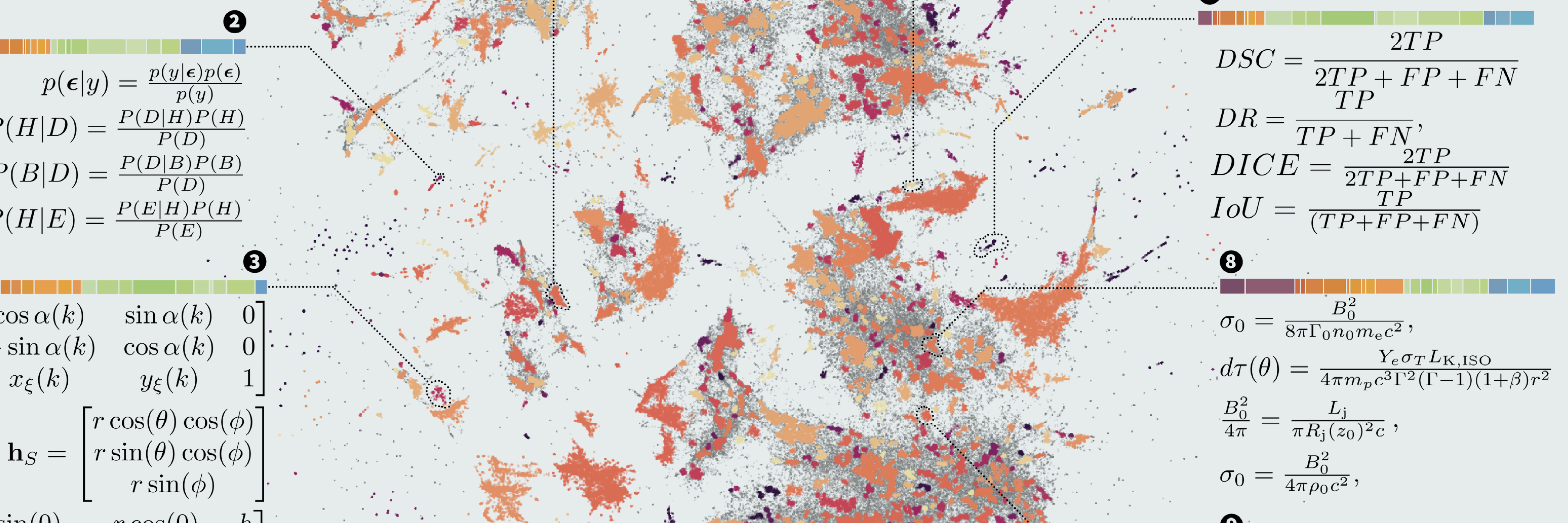
A new paper, in which I try to figure out how common the interdisciplinary distribution of computational templates is in the sciences. It turns out: Very common! –
Read more...
Sally is simply \\ perceiving a pineapple \\ in the normal way. –
Read more...
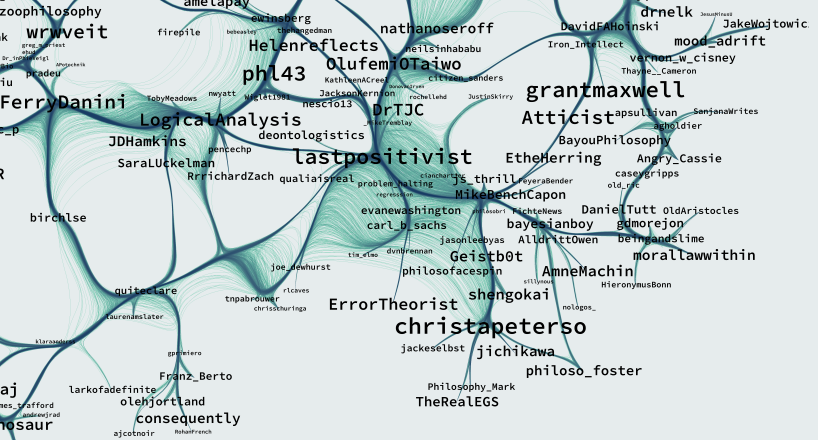
A map of the network of academic-philosophy-twitter. –
Read more...
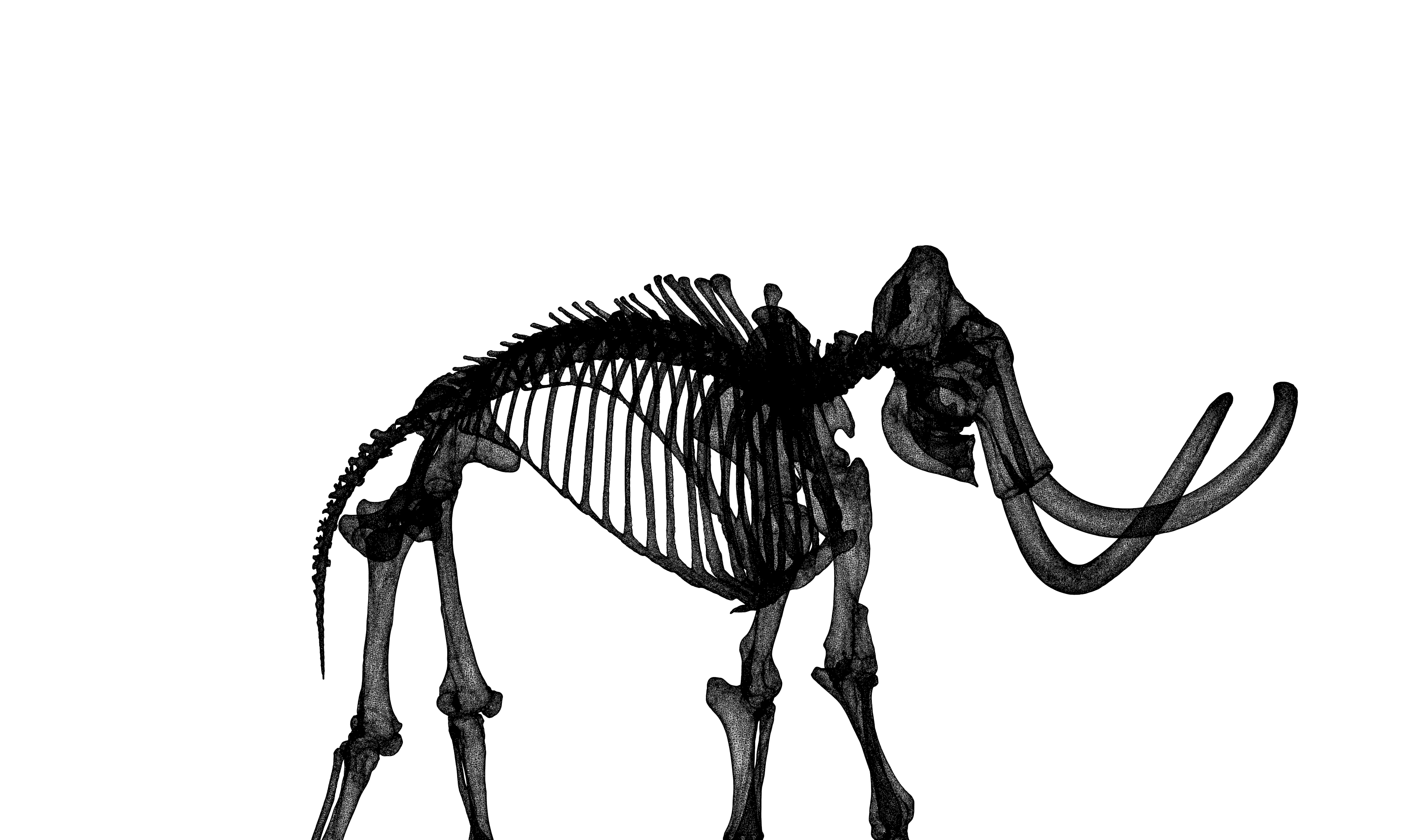
An illustration of non-linear dimensionality-reduction using prehistoric pointclouds. –
Read more...